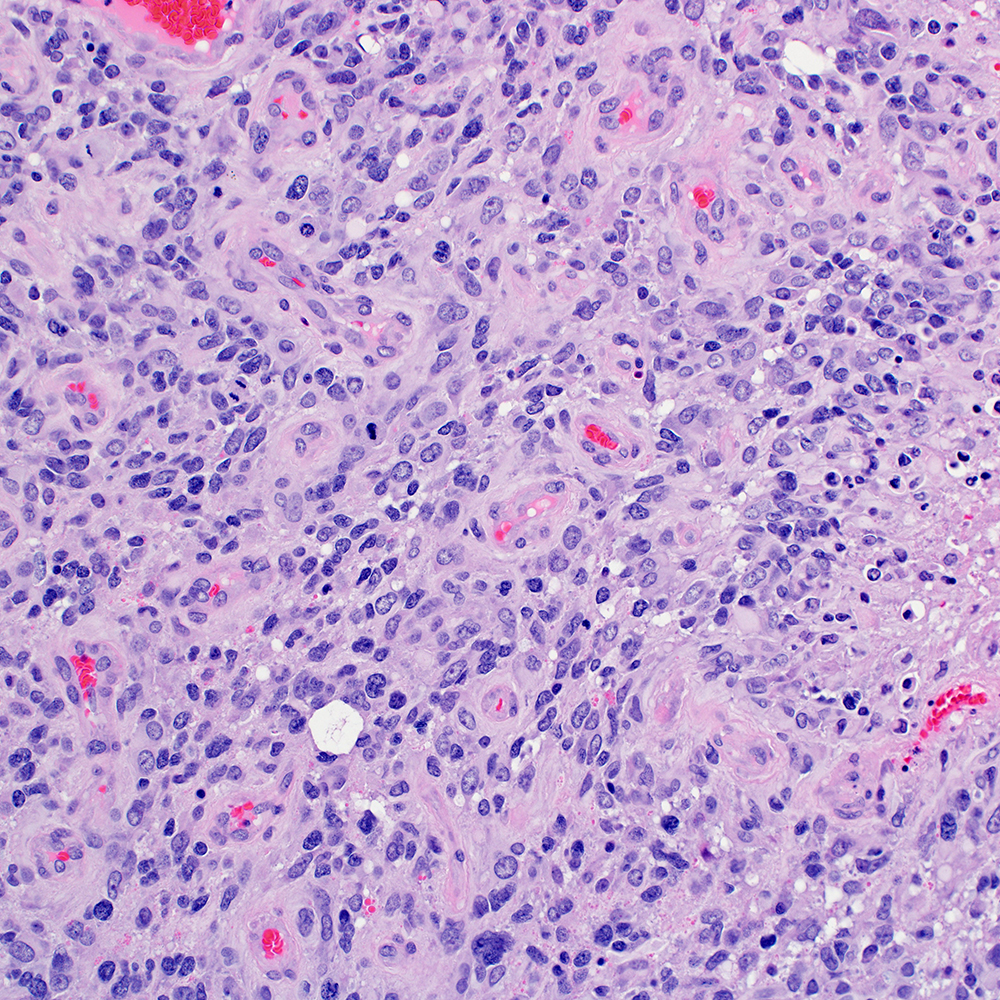
Model Systems
Sharing patient-derived xenograft models and data generated at St. Jude to stimulate basic research and speed translation to the clinic.
About Model Systems
Research programs at St. Jude generate a wide range of pediatric patient-derived xenografts (PDX) and patient-derived orthotopic xenograft (PDOX/O-PDX) models. As part of the St. Jude Cloud Initiative, these resources provide both in vivo and in vitro systems to study mechanisms of disease origin, tumor progressions, and preclinical testing.
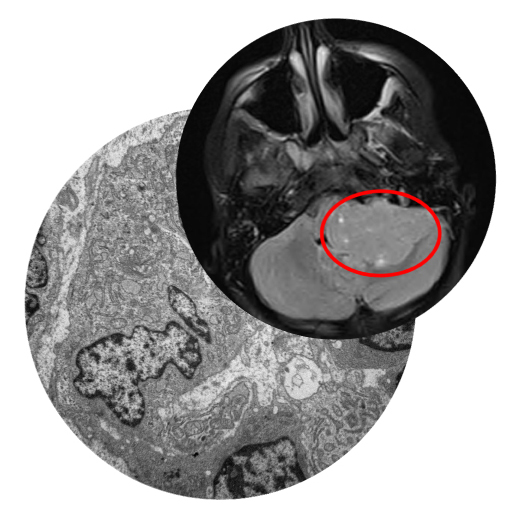
The Childhood Solid Tumor Network (CSTN), Pediatric Brain Tumor Portal (PBTP), and Public Resource of Patient-derived and Expanded Leukemias (PROPEL) provide hundreds of PDX and PDOX/O-PDX models consisting of over 40 diagnoses, including high grade glioma, medulloblastoma, rhabdomyosarcoma, neuroblastoma, over 20 subtypes of leukemia, and rare solid tumors. These resources provide xenograft cells for propagation and preclinical testing and curated data gathered when characterizing the models. Raw sequence files are available through our Genomics Platform.
In addition to making the data freely available, St. Jude’s ongoing effort to accelerate and advance cures of catastrophic pediatric diseases, these model systems provide an easy pathway for requesting cryopreserved cells for propagation and preclinical testing. These resources also include FFPE tissue sections, fresh frozen tissue, pharmacokinetic data, protocols used to generate the models and data from them.
Featured Study Programs
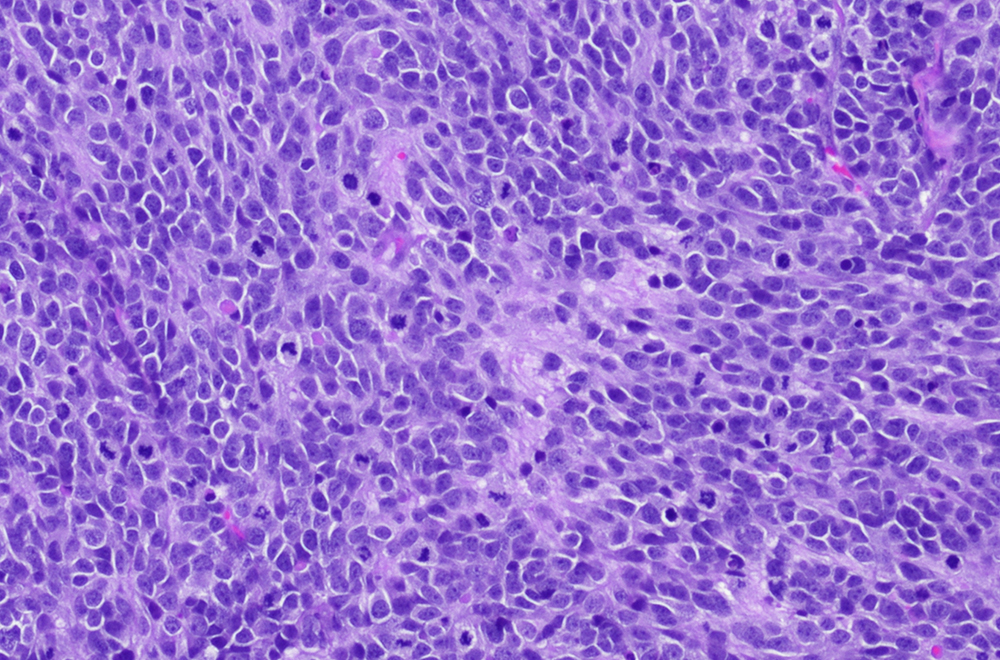
Childhood Solid Tumor Network
The CSTN Data Portal provides access to orthotopic patient-derived xenograft models of childhood solid tumors and data.
Explore CSTN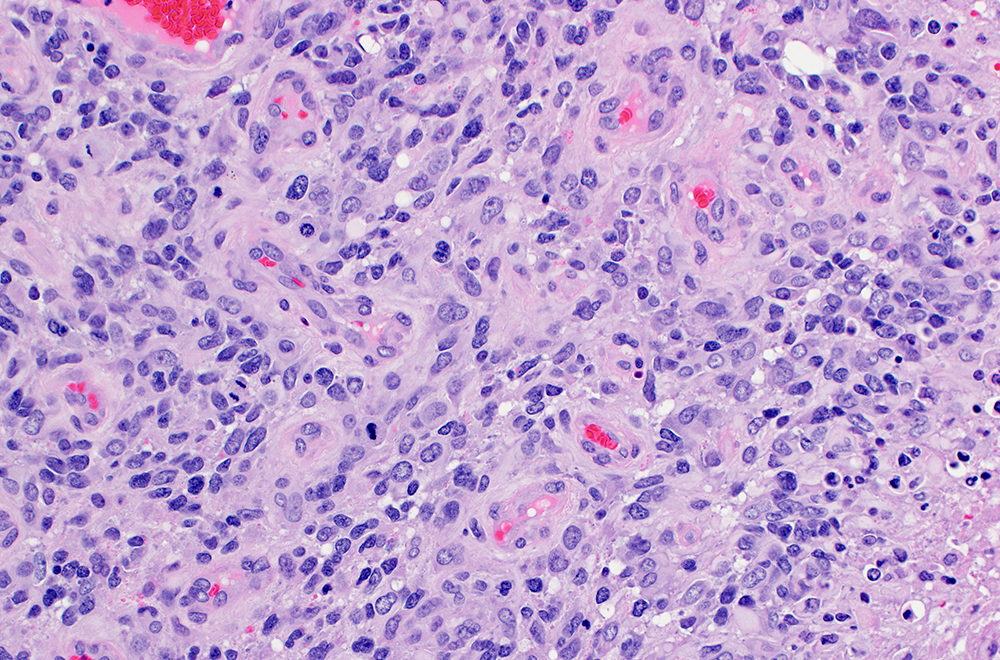
Pediatric Brain Tumor Portal
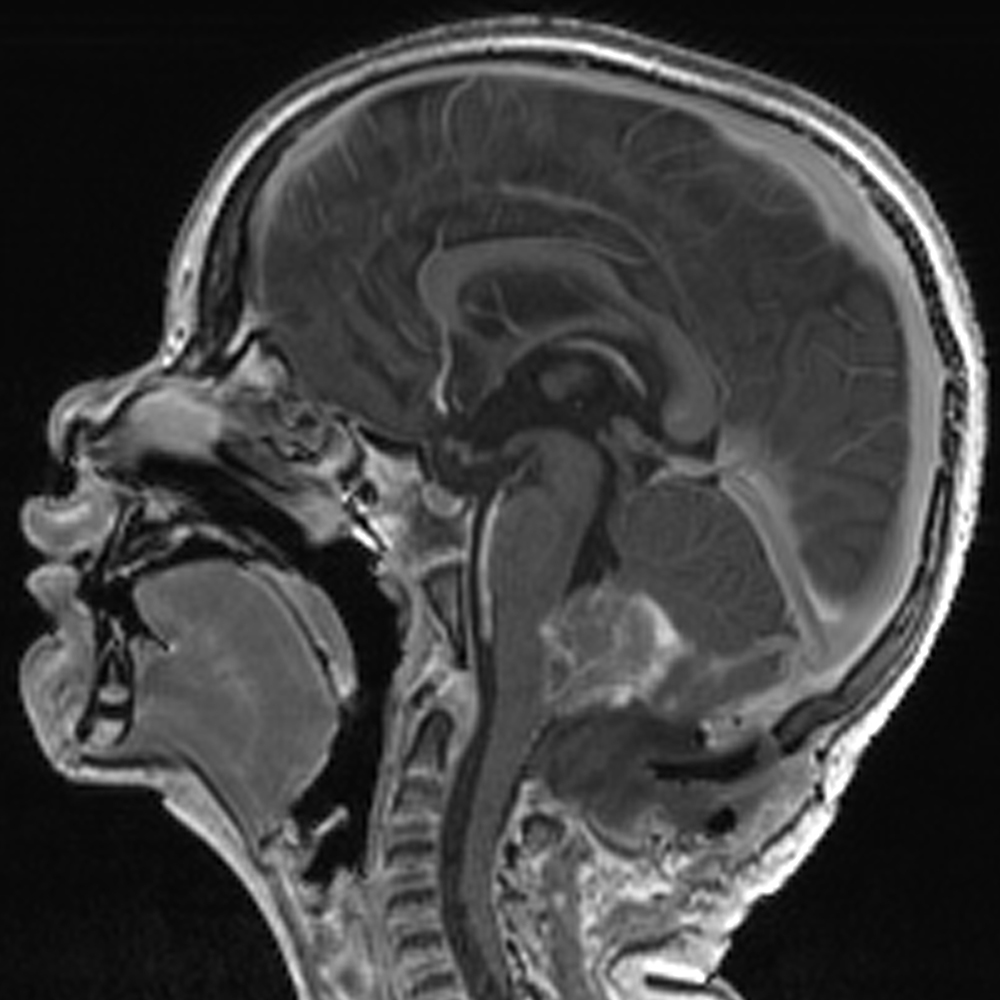
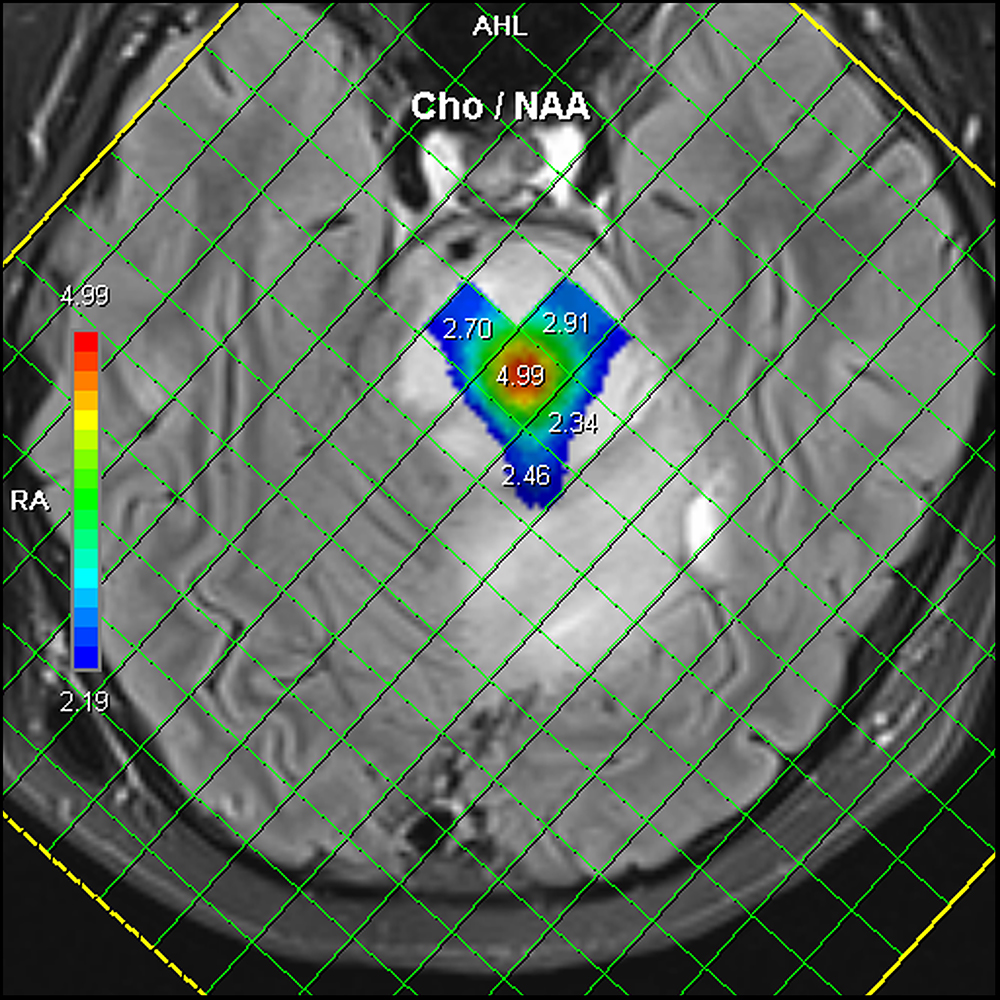
The PBTP Data Portal provides access to molecular characterization and orthotopic patient-derived xenograft models of childhood pediatric brain tumors and matched cell lines from diverse groups of pediatric high grade gliomas.
Explore PBTP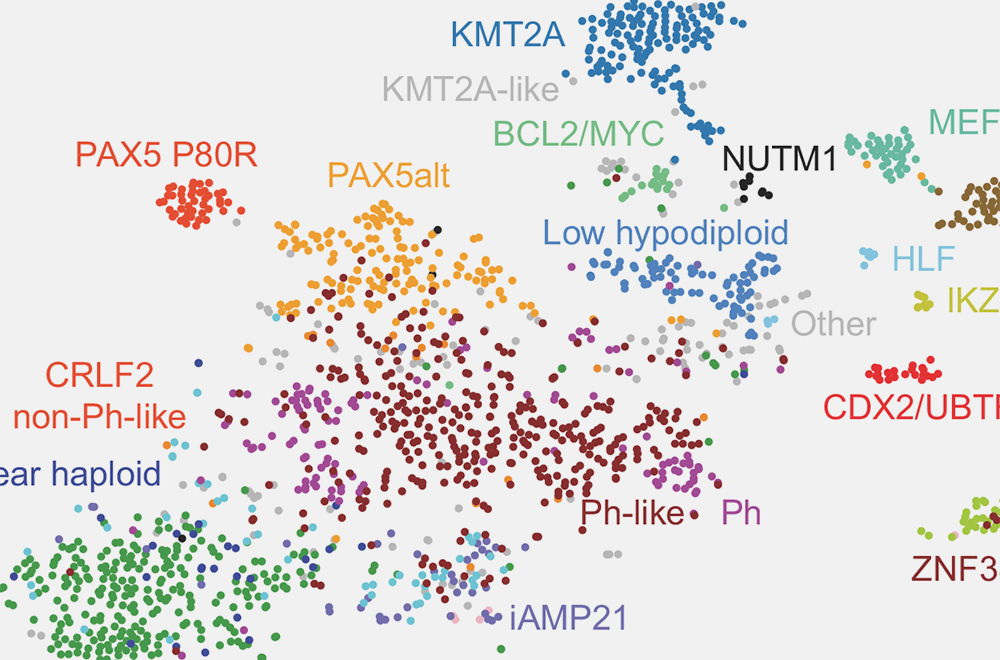
Public Resource of Patient-Derived and Expanded Leukemias
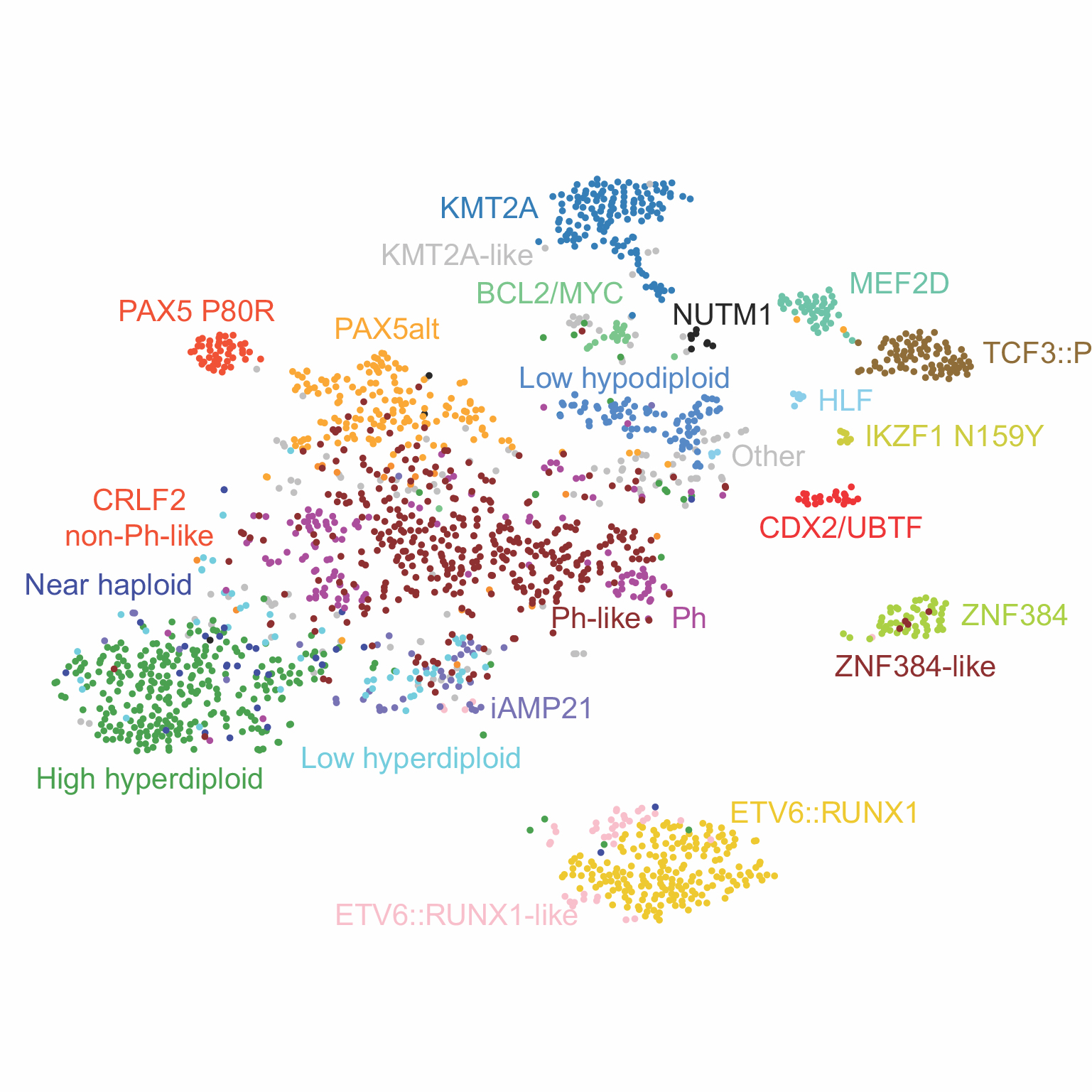
The PROPEL data portal is one of the world's largest repositories of PDXs for adult and pediatric leukemias.
Explore PROPELHigh-Quality Data
Our raw genomics data as well as data from multi-omics provides a rich resource for the research community to find the appropriate models to advance the knowledge and therapeutic solutions to pediatric cancer.
Cohort-Level Data
Explore curated data for somatic mutations, differential gene expression, DNA methylation classification, disease subgrouping, and chemical sensitivity data for patient and/or matched patient samples as well as PDOX/O-PDX-derived cell lines.
Sample-Level Data
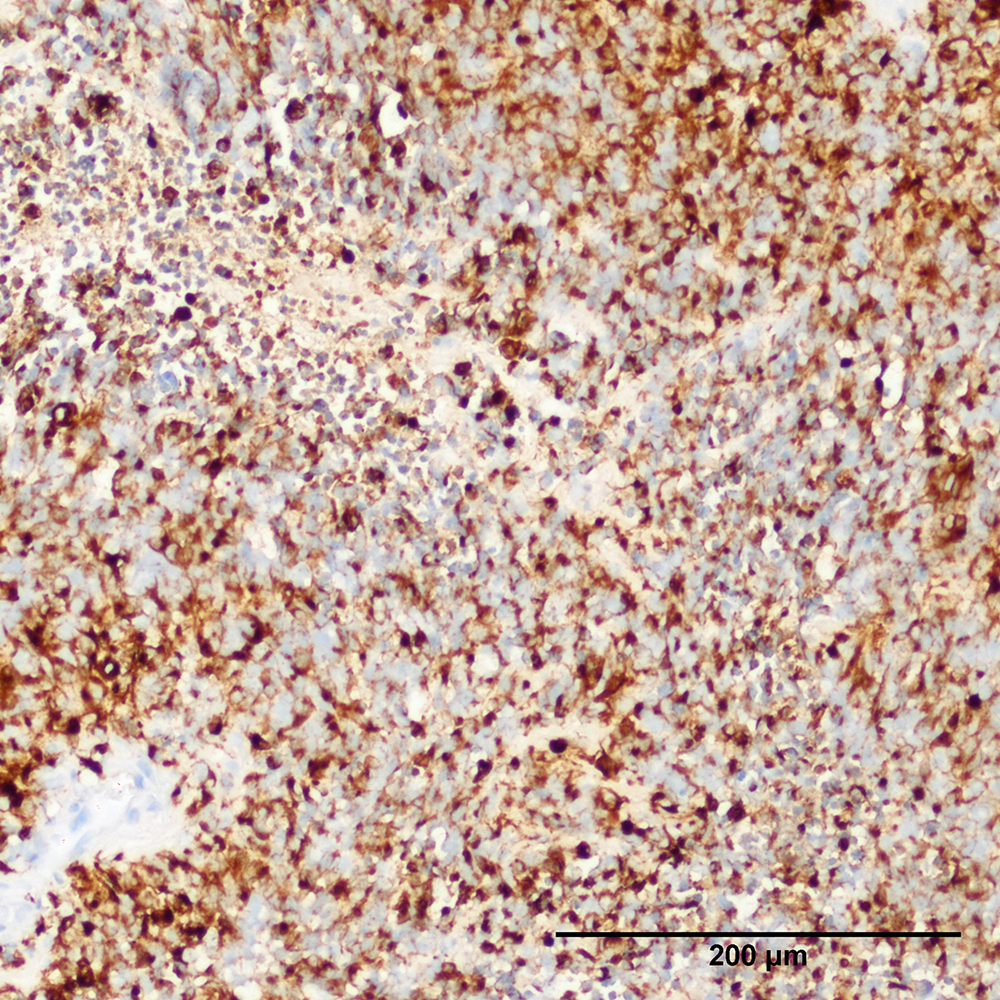
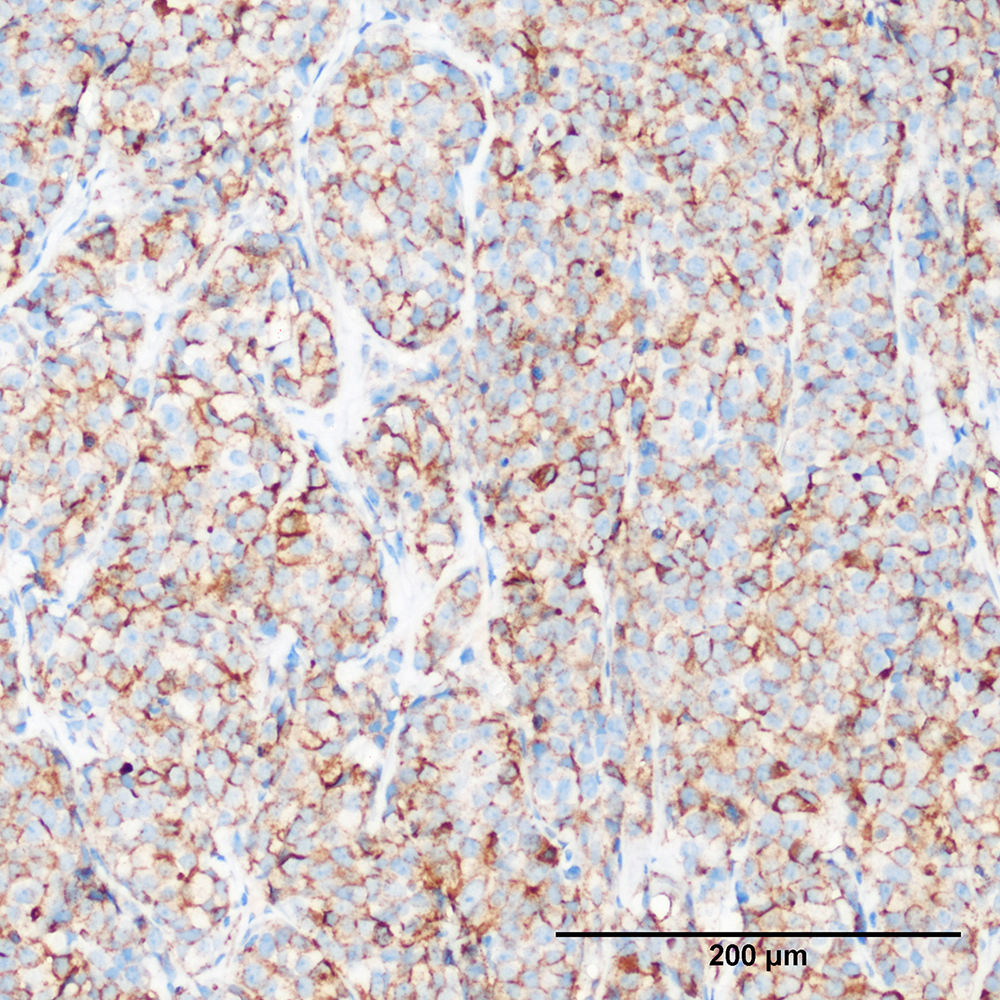
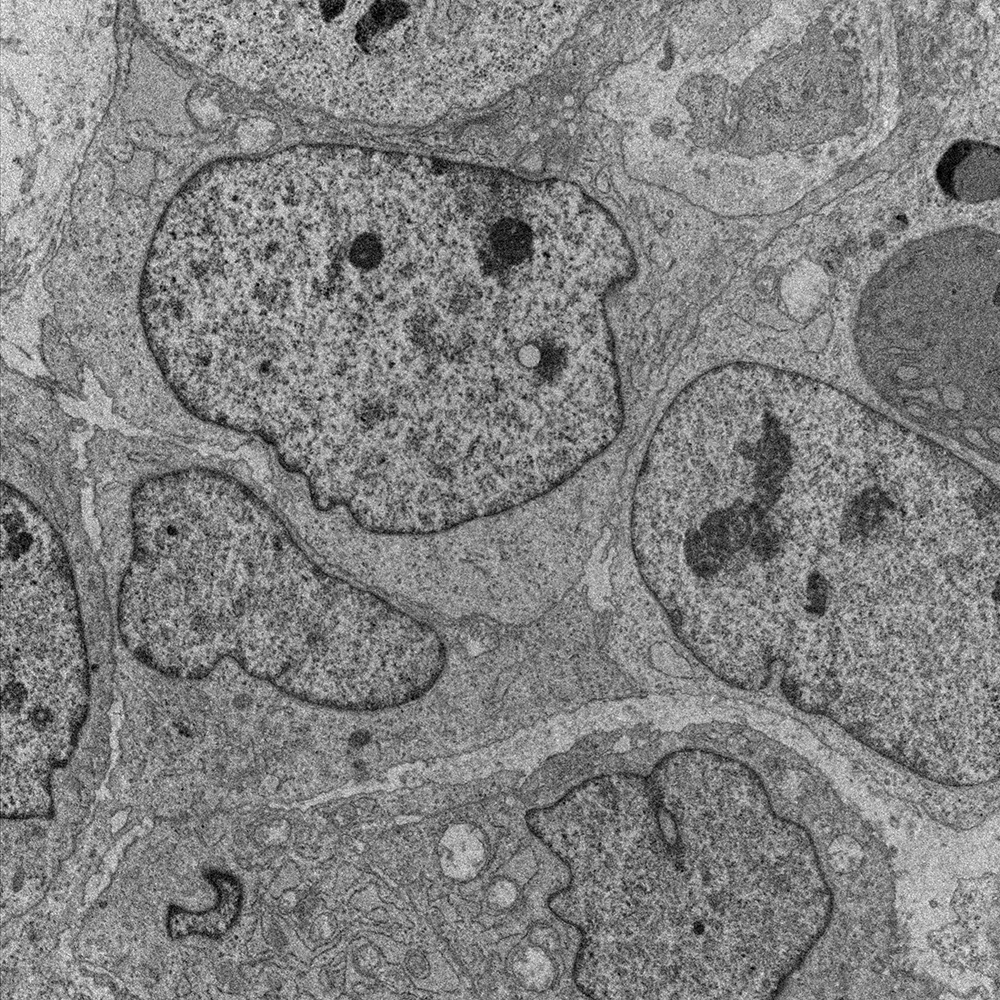
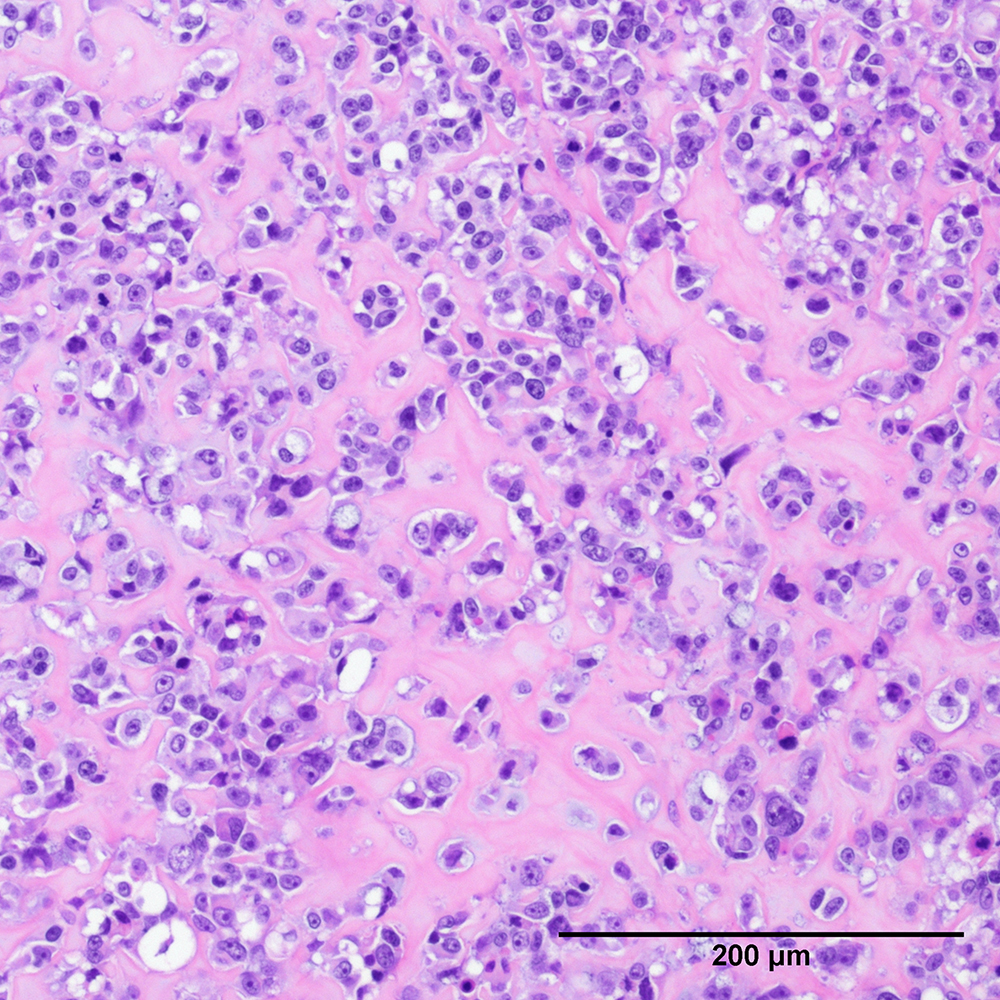
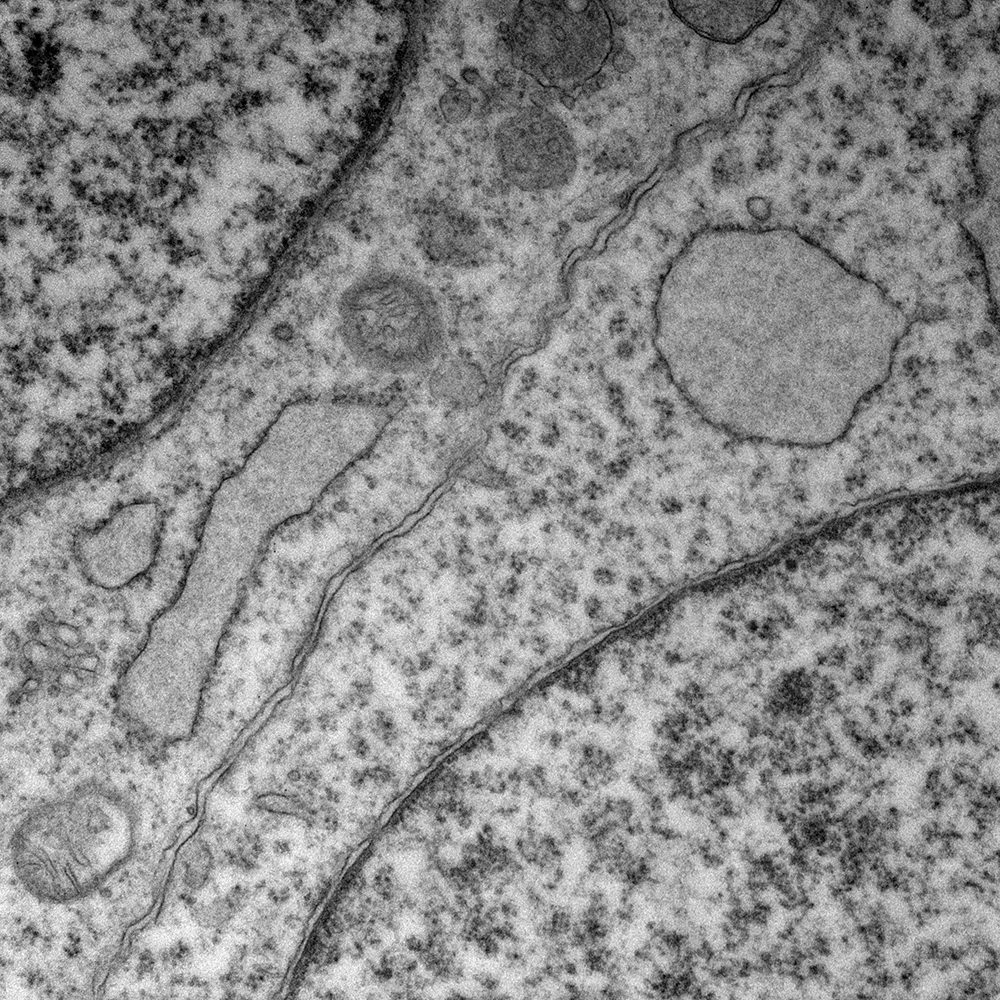
Discover comparative histology, electron microscopy, copy-number variations, and Fluorescence in situ hybridization (FISH) of our samples and data.
Gene-Level Data
Search genes of interest, generate custom oncoprints, and explore single nucleotide and structural variants.
St. Jude Cloud
St. Jude Cloud, in partnership with Microsoft and DNAnexus, aims to share data and analysis resources to the global research community. Our goal is to support, advance, and accelerate cures for pediatric cancer and other pediatric catastrophic diseases, like sickle cell disease.
Dive into our other applications such as the Pediatric Cancer Knowledgebase (PeCan) or the visualization tools ProteinPaint, GenomePaint, and Disco. These resources are available for free with no obligation to collaborate.